![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/3-Table2-1.png)
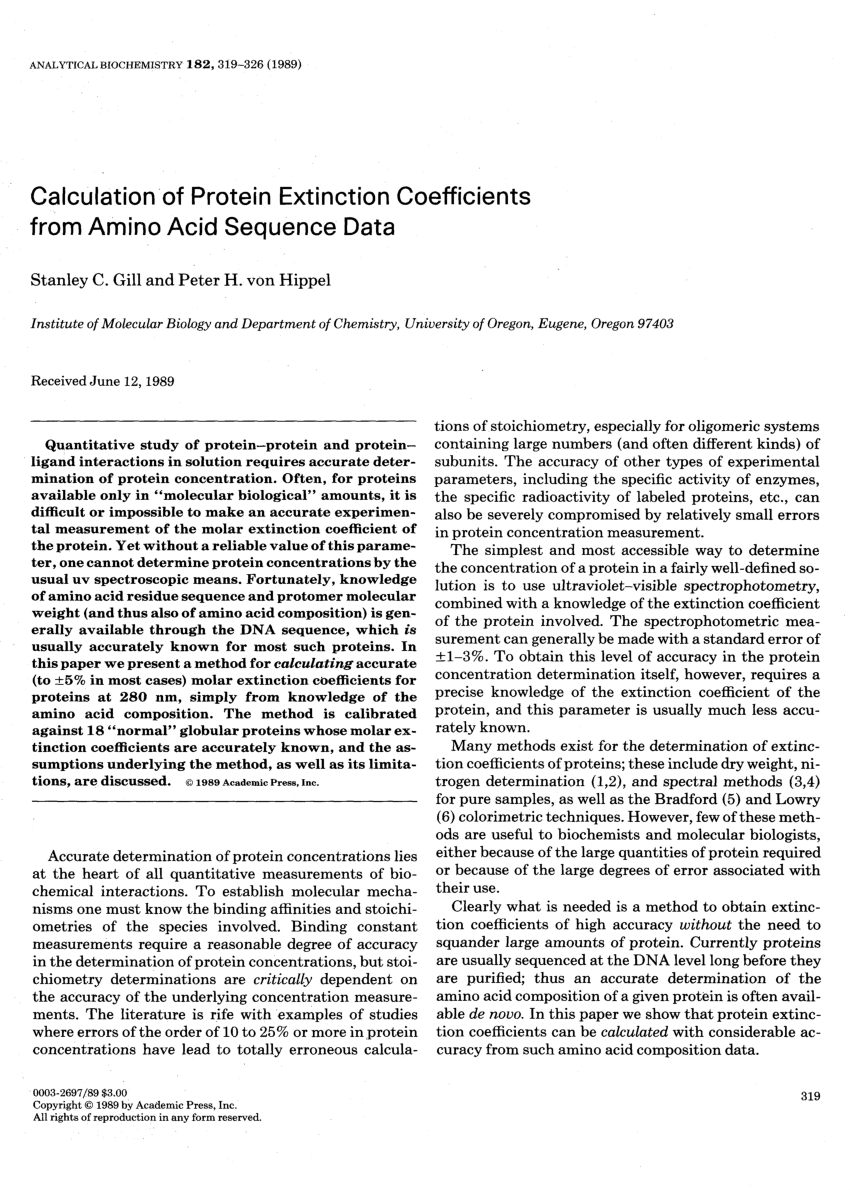
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar
Complex folding and misfolding effects of deer-specific amino acid substitutions in the β2-α2 loop of murine prion protein | Scientific Reports
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/4-Table3-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar

Artificial intelligence and machine learning for protein toxicity prediction using proteomics data - Vishnoi - 2020 - Chemical Biology & Drug Design - Wiley Online Library
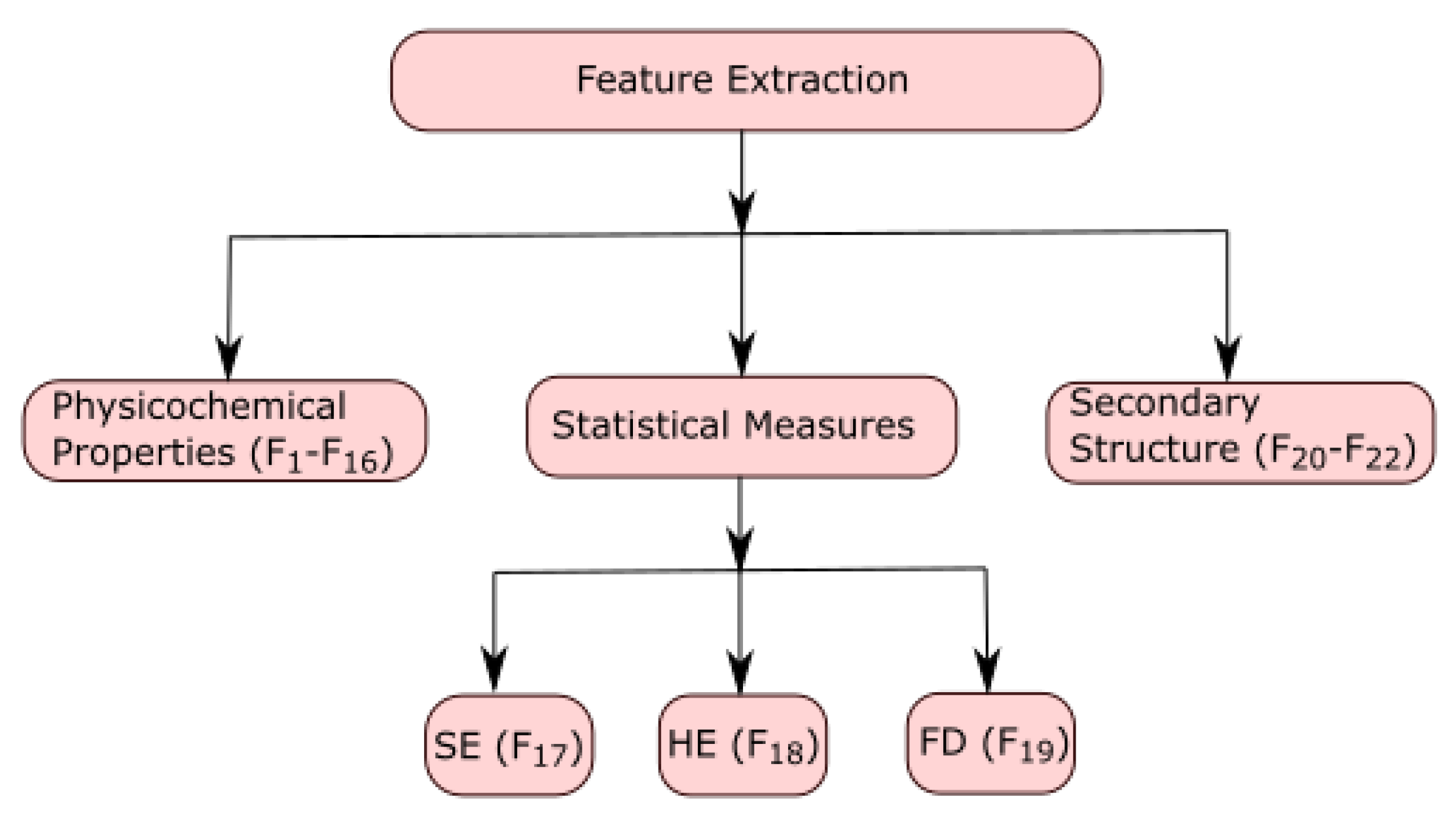
Mathematics | Free Full-Text | Unsupervised Learning for Feature Representation Using Spatial Distribution of Amino Acids in Aldehyde Dehydrogenase (ALDH2) Protein Sequences
Engrailed and Hox homeodomain proteins contain a related Pbx interaction motif that recognizes a common structure present in Pbx. - Abstract - Europe PMC
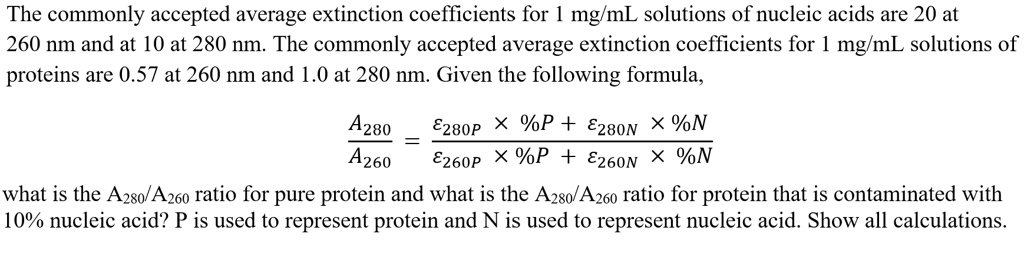

SOLVED: The commonly accepted average extinction coefficients for mg/mL solutions of nucleic acids are 20 at 260 nm and at 10 at 280 nm The commonly accepted average extinction coefficients for mglmL

Precise determination of protein extinction coefficients under native and denaturing conditions using SV-AUC | SpringerLink

Photometric Quantification of Proteins in Aqueous Solutions via UV-Vis Spectroscopy - Eppendorf Handling Solutions

A method to identify and quantify the complete peptide composition in protein hydrolysates - ScienceDirect

Prediction of Molar Extinction Coefficients of Proteins and Peptides Using UV Absorption of the Constituent Amino Acids at 214 nm To Enable Quantitative Reverse Phase High-Performance Liquid Chromatography−Mass Spectrometry Analysis | Journal